
WHAT PONTEGRA PROVIDES
Build repeatable cohort-to-feature pipelines from FHIR and OMOP on Apache Spark
Pontegra helps life sciences and real-world evidence teams turn longitudinal clinical data into analysis-ready datasets that are fast to build, repeatable to run, and scalable to production volumes.
Powered by: Studyfyr – The Governed Clinical Research Toolkit and Platform on FHIR and OMOP.
WHAT YOU CAN ACHIEVE
Scalable Science, Reproducible Results
Accelerate time-to-dataset
Move from data access to analysis-ready tables without one-off flattening scripts.
Make studies reproducible
Define cohort logic, sampling rules, and feature definitions as rerunnable pipelines.
Scale to real-world volumes
Run consistent logic across millions of patients and large event histories with Spark-native execution.
Support iterative study development
Start with a scoped sample for early validation, then scale to full runs.
Why Studyfyr for RWE Processing
Eliminating the “Data Tax” on Clinical Research
Most RWE teams lose time on:
- normalizing longitudinal events,
- aligning timelines to index dates,
- generating covariates and outcomes consistently,
- keeping refresh cycles stable as data grows.
Studyfyr reduces this burden with:
- FHIR-native query semantics (FHIR search and FHIRPath), also adapted for OMOP,
- FHIR/OMOP compatible cohort and dataset workflows,
- Spark scale-out execution,
- reusable building blocks for cohorts, sampling, features, and outcomes.
HOW IT CAN BE DONE
An Automated Journey from Raw Data to Insight
0.
Select data source
- FHIR API (server)
- FHIR lake (NDJSON/Parquet/Delta)
- OMOP database/lake extracts
1.
Define cohort logic
- Entry, exit criteria
- Choose eligibility period(s) (for example first diagnosis, first prescription, all medication exposures)
- Inclusion and exclusion rules
2.
Define sampling strategy
- Periodic sampling (monthly, quarterly)
- Event-aligned sampling (for example pre/post treatment or admissions/discharges)
- Outcome-aligned sampling (for example 1,2, … hours before complication in ICU)
- Rolling windows such as “since index” or “since last timepoint”
3.
Compute features and endpoints
- Covariates (labs, vitals, diagnoses, medications, utilization)
- Endpoints and outcomes (events, counts, time-to-event derivations)
- Feature enumeration over temporal windows (tumbling, extending, session, etc) and aggregations (avg, max, min, last, count)
4.
Produce analysis-ready outputs
- Relational and feature datasets for HEOR, epidemiology, and AI workflows
- Reproducible derived tables for downstream R/Python/ML pipelines
5.
Optional: Promote / Publish to organizational level
- Promote approved pipelines/artifacts for reuse
- Publish periodic dataset snapshots for governed consumption
Typical RWE Pipeline Patterns
Cohort feasibility and characterization
- Define cohort logic once
- Generate counts and baseline characteristics
- Iterate quickly with scoped runs
Confounders and covariates
- Compute time-windowed features before index date
- Derive exposure, comorbidity, and laboratory summary variables
Endpoints and outcomes
- Generate event tables and derived outcomes
- Build longitudinal patient-by-timepoint modeling tables
Refreshable production pipelines
- Rerun with newly arrived data
- Maintain stable definitions for repeated analysis cycles
where ıt runs
Run Wherever Your Data Lives
Runs wherever Spark runs, including Kubernetes, YARN, Databricks, EMR, and on-prem clusters.
You can use Studyfyr SDKs in notebook environments using:
- Pontegra CRT (open source): Spark/Scala SDK for clinical data processing
- Pontegra Studyfyr (enterprise): governed execution layer with
- managed service API and control plane UI
- Python SDK (thin client) for workflow execution
Example Notebook Snippets (SDK)
Example 1: Cohort pipeline

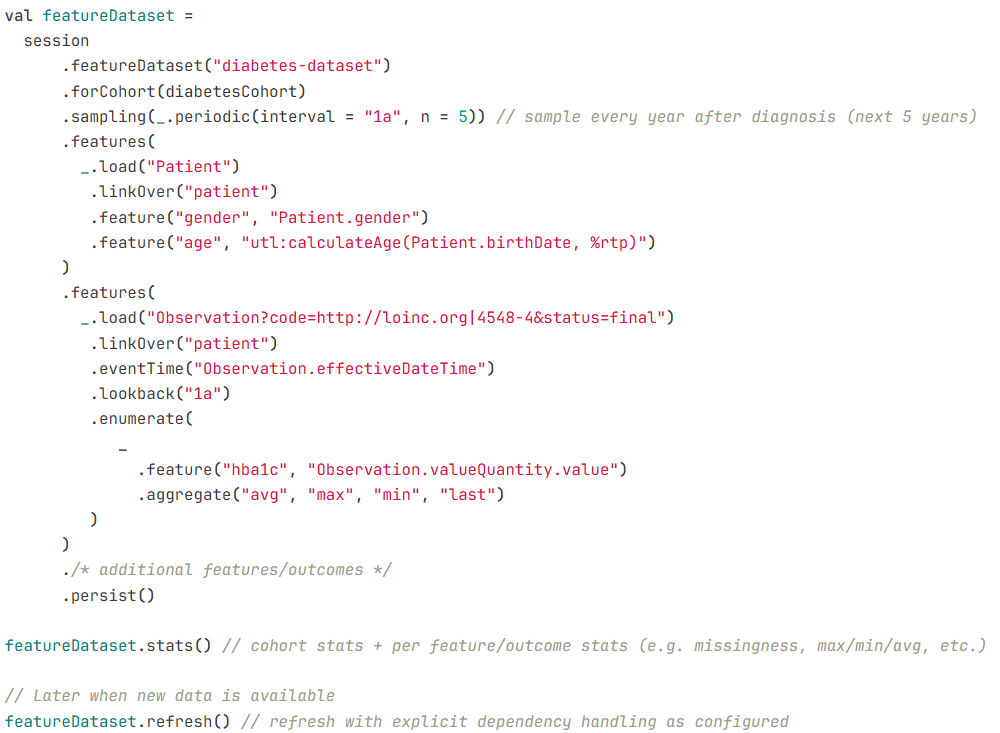
Example 2: Feature dataset pipeline
Governance by Default
Even in processing-first deployments, Studyfyr keeps governance signals built into execution.
For EHDS/SPE/TRE governance details, see the EHDS-aligned solution page.
- Run manifests capture inputs, definitions, versions, outputs, and timing.
- Metadata-first audit records support operational traceability (no PHI in audit by default).
- Controlled publish/export patterns enforce review before organizational or external release.
Frequently asked questions
Does this support both FHIR and OMOP?
Yes. Pipelines can run on FHIR and OMOP sources, including API, database, and lake-based patterns.
Do we need to change our R/Python modeling stack?
No. Studyfyr/CRT produces analysis-ready datasets for your existing R/Python/ML workflows.
Where does it run?
Wherever Spark runs (for example Databricks, EMR, Kubernetes, YARN, and on-prem clusters).
How are reruns and updates handled?
Pipeline runs are reproducible by design, with manifest-backed execution and refresh/recompute lifecycle patterns.
Can this operate in governed environments (SPE/TRE)?
Yes. Controlled execution and output review patterns support secure secondary-use workflows.
Pilot Delivery Plan
A typical pilot focuses on one therapeutic area and delivers the following.
- connection to one approved source (FHIR or OMOP)
- one cohort definition with entry/exit/eligibility logic
- one sampling strategy (periodic or event-aligned)
- two to three analysis-ready dataset tables
- run documentation (definitions, parameters, outputs, manifest references)
SEE YOUR OPTIONS
Packages
Available now
Design Partner Track
- Best for: Organizations co-shaping early implementation priorities.
- Timeline: Ongoing
- Typical output: Use-case backlog, phased rollout plan, early access alignment.
Planned June 2026 *
Secondary Use Starter
- Best for: Teams ready to implement a first governed cohort-to-dataset pipeline.
- Timeline: 4-6 weeks
- Typical scope: One cohort definition, one dataset specification, pipeline implementation, run manifest/audit outputs, handover.
Planned October 2026 *
Secondary Use Scale-Up
- Best for: Multi-study rollout and recurring governed operations with Studyfyr.
- Timeline: 3-6 months
- Typical scope: Additional cohorts/datasets, scheduled refresh/recompute cycles, operating model hardening.
* Planned dates may be adjusted.